Unusual Ways to Lose a Y Chromosome and Survive with Changed Autosomes: a Story of Mole Voles Ellobius (Mammalia, Rodentia)
Irina Bakloushinskaya 1,* ![]() , Sergey Matveevsky 2
, Sergey Matveevsky 2 ![]()
- Koltzov Institute of Developmental Biology, Russian Academy of Sciences, Moscow 119334, Russia
- Vavilov Institute of General Genetics, Russian Academy of Sciences, Moscow 119991, Russia
* Correspondence: Irina Bakloushinskaya![]()
Received: May 16, 2018 | Accepted: July 13, 2018 | Published: July 23, 2018
OBM Genetics 2018, Volume 2, Issue 3 doi:10.21926/obm.genet.1803023
Academic Editors: Miodrag Stojkovic and Darren Griffin
Special Issue: Reproductive Genetics
Recommended citation: Bakloushinskaya I, Matveevsky S. Unusual Ways to Lose a Y Chromosome and Survive with Changed Autosomes: a Story of Mole Voles Ellobius (Mammalia, Rodentia). OBM Genetics 2018;2(3):023; doi:10.21926/obm.genet.1803023.
© 2018 by the authors. This is an open access article distributed under the conditions of the Creative Commons by Attribution License, which permits unrestricted use, distribution, and reproduction in any medium or format, provided the original work is correctly cited.
Abstract
Species of mole voles Ellobius demonstrate a broad variation in sex chromosomes and autosomes, which is unique among mammals. In four species, a Y chromosome was lost, and X0 or XX sex chromosomes in both sexes were obtained. The key testis-determining Sry (Sex-determining Region on Y) gene is absent in these species, and the regulation of its target, the Sox9 (SRY -box 9) gene, is questionable due to deletion in the key enhancer. In a single species, E. fuscocapillus, with routine XX-XY, the same deletion is present alongside fragments of Sry in the female genome. Presumably, a Y chromosome was lost twice in two phylogenetic lineages of mole rats; before the event, a few male-specific genes escaped on X chromosomes. Translocations of Y chromosome fragments were made independently, resulting in different changes in species without a Y chromosome and the presence of the Y-linked Sry gene in females of E. fuscocapillus, a species retaining the Y chromosome. One more exceptional phenomenon is high autosomal variability in E. tancrei. This species might be used as an exclusive model for studying meiotic mechanisms providing balanced gametes in complex heterozygous hybrids. Sterility is the only destiny for hybrids, whose parents carry Robertsonian translocations with partial homology. Contrary to that, E. tancrei possess different Robertsonian translocations and successfully overcome the hybrid incompatibility. Here, we overview the research to date of sex determination and meiosis in Ellobius.
Keywords
Sex determination; meiosis; sex chromosomes; Robertsonian translocations
1. Introduction
Gene interaction during early development determines the development of gonads, producing gametes in a distinct way in males and females. Meiosis is essential to provide a precise transmission of parental genetic information to the next generations, which is a base for effective reproduction and evolutionary success. The evolutionary plasticity of a primary switch of sex determination in vertebrates, such as genetic, epigenetic, temperature-dependence or other types of elusive nature, remains poorly understood [1,2,3]. Three modern infraclasses of mammals (monotremes, marsupials and placental) have significant differences in their sex chromosomes. Monotremes, the most diverged mammalian group, obtained multiple sex chromosomes — 10 elements in platypus males and 9 in echidna males [4,5]. Marsupials and placental mammals separated from monotremes approx. 166 million years ago [6], and evolved with sex chromosomes XX-XY, the gene content and structure of sex chromosomes appeared to be different in the groups. Evolution of their heteromorphic sex chromosomes, as suggested due to molecular studies of a Y chromosome, started approximately 180 million years ago in an ancestor of Eutheria [7].
A Y chromosome evolution might be initiated by the emergence of a specific testis determining factor and decreasing recombination between originally homomorphic chromosomes. Subsequently, these chromosomes evolved into heteromorphic chromosomes associated with sex [8,9]. The suppression of recombination developed as a result of numerous chromosome changes, mostly inversions, creating at least four evolutionary strata at sex chromosomes [10]. The schematic network of sex determination should include several components: a primary genetic signal, the key gene, which is a target to this primary signal, and a gene switching between two alternative programmes of development [11].
Sex determination is a strictly scheduled process when the fine-tuned gene expression levels are provided. In placental mammals the gene SRY (Sex-determining Region on Y) is a key factor that determines the formation of testes (the Testis Determining Factor, TDF) and locates on the Y chromosome [12,13,14]. The prime role of the SRY is to activate the SOX9 (SRY -box 9), which should later (after 11.5 days post coitum (dpc) in mice) activate other genes in the male pathway, such as fibroblast growth factor 9 (FGF9) and fibroblast growth factor receptor 2 (FGFR2). The FGF9 and FGFR2 genes normally block Wnt4 (Wnt family member 4) in males. In a case of down regulation of FGF9, Wnt4 blocks SOX9 and starts the female pathway by activating β-catenin, forkhead box L2 (FOXL2) and others. The idea that female sex is programmed by default and a specific male-determining factor was originated to override it has prevailed since the Sry gene was described (for review see [1]). However, the process of sex determination is apparently more complex. First, there is data on the epigenetic regulation of Sry expression and Sertoli cells differentiation, such as DNA methylation and histone modifications, for example, by the histone demethylase JMJD1A (Jumonji domain containing 1A) and others [15,16,17]. The ovarian development in mice starts later, at 12.5 dpc, and now it is clear that a ‘default’ mechanism is not efficient. The ovarian pathway is studied intensively, with a main accent on the RSPO1 (R-spondin1) gene [18]. Whereas male (testes) development depends on a single pathway of SOX9 regulation, the female (ovaries) development is controlled by at least two ways of SOX9 suppression [19]. The intensive studies of signalling pathways involved in mammalian sex determination revealed a tremendous amount of regulation events, including epigenetic ones [3,16,17]. One of the most amazing features of sex determination is a possibility of sex reversal in adult mammals. That means a transformation of testes to ovaries and vice versa and entails a life-long struggle against an alternative sex determining pathway by the repression of several key genes [20,21]. The early mammalian gonad contains bipotential precursor cells for steroid-secreting cells and supporting cells, which sustain and nourish germ cells. In male gonads, supporting cells can develop into Sertoli cells; in ovaries, to specific follicle (granulosa) cells. Sex determination and a reverse process both depend on the state of supportive cells, which should achieve a threshold in early development for getting ‘proper’ gonads, but even in adulthood, these supportive cells should keep a status quo. It was shown that in adult mice, an ablation of Foxl2 in granulosa cells leads to the Sox9 expression, which induces the transdifferentiation of granulosa cells to Sertoli cells [20]. In contrast, the deletion of the Dmrt1 (doublesex and mab-3 related transcription factor 1) in testes results in the Foxl2 upregulation and transformation of testicular Sertoli cells to ovarian granulosa cells [21]. Sox9 and Sox8 (SRY-box 8) together with Dmrt1 protect the adult testis from male-to-female genetic reprogramming and complete degeneration [22].
The Sry gene is present in the marsupial male genome and probably has a function as a sex determination gene [23,24]. There are also exceptions to the rule in placentals: several rodents have lost a Y chromosome and a primary sex determining gene Sry, such as several species of mole voles Ellobius [25] and spiny rats Tokudaia [26].
Our study was focused on the analysis of chromosome changes in mole-voles Ellobius from different points of view, with an emphasis on the meiosis and, especially, the behaviour of sex chromosomes, specific transmission of Robertsonian translocations between generations and variability of genes involved in sex determination and spermatogenesis. The primary hypothesis was that these phenomena are independent, and we tried to separately analyse the evolution of sex determination, genes and chromosomes, as well as autosomal variability in species with two X chromosomes in males and females. However, as we obtain more data, we have come to a conclusion that the evolution of sex chromosomes can probably play a leading role. Changes in sex chromosomes may enforce a genomic conflict, hybrid incompatibility, and, as an evolutionary output, generate reproductive barriers leading to species divergence [27]. Such ways of evolution might be favourable, even for such phylogenetic lineages, as mammalian infraclasses [28]. In this review, we aim to highlight the evolutionary history and uniqueness of sex determination without a Y chromosome and the elusiveness of primary sex determining genes in mole voles Ellobius alongside their special meiotic behaviour of sex chromosomes and variable autosomes.
2. Ellobius Species and Their Sex Chromosomes Variety
Mole voles Ellobius were described by Pallas in the XVIII century as Mus talpinus Pallas, 1770 [29], and remained poorly studied for over a hundred years, perhaps because of their secretive subterranean lifestyle. Fisher, in 1814, affiliated the species to a new genus Ellobius [30], which contained from two up to five species in a different taxonomy system [31]. R. Matthey, a famous Swiss cytogeneticist, revealed a unique mammalian odd diploid chromosomal number in E. lutescens (2n=17), and a single X chromosome in males and females [32]. An ordinary mammalian sex chromosome constitution XY♂ / XX♀ was recognized in the southern mole vole E. fuscocapillus (2n = 36) [33]. Three morphologically indistinguishable species lack a Y chromosome: the northern mole vole E. talpinus (diploid number 2n = 54, the total number of chromosome arms NF = 54), the eastern mole vole E. tancrei (2n = 54-30, NF = 56) and the Alai mole vole E. alaicus (2n = 52, NF = 56); all these species have two X chromosomes in females and males [33,34,35,36]. It was assumed that the whole or partial Y chromosome was translocated to the X chromosome or autosomes, but did not disappeared entirely, because males develop typical testes, and the sex ratio in all mole vole species is normal: approx. 50% of newborns are males, and 50% are females. The structures of autosomes and sex chromosomes were studied by chromosome painting of 4 species lacking Y chromosomes; the X chromosome is typical for rodents, and no parts of the Y chromosome were revealed [37,38]. To date, only E. fuscocapillus was not studied by molecular cytogenetic methods. The chromosome painting data made it possible to reconstruct an ancestor karyotype for mole voles and evaluate changes in comparison with the Arvicolinae ancestor karyotype [37,39].
The discovery of an odd chromosomal number in an E. lutescens karyotype arose a question of evolutionary history and functionality of a single X chromosome. How were a Y and the second X chromosome lost? How did the identical X chromosomes in males and females provide different sex determination? First, a hypothesis of fusions of two Xs in females and X with Y in males was declared [40,41]. Nevertheless, studying meiosis [42,43,44,45] and applying competitive fluorescence in situ hybridization with genomic DNA probes (comparative genomic hybridization, CGH) disproved the hypothesis [46]. The 9th chromosome was proven as the X because the gene G6pd (glucose-6-phosphate dehydrogenase), typical for Xs, was detected on it [44]. Later, a comparative chromosome painting proved the assumption; the clear signal was distinguished on the 9th chromosome with X chromosome probes of several mammalian species, such as humans, mice, rats, Microtus agrestis, and Mesocricetus auratus [37], and no signal was detected for any autosomal probes. It is important that neither differences for male and female chromosomal sets, nor the signal for the Y chromosome probes were revealed. The same results were obtained for species with two X chromosomes in males and females of Е. talpinus and E. tancrei [38]. These results proved the total loss of a Y chromosome, but chromosome painting has a restriction because a fluorescent signal might be unrecognizable in the case of a small fragment, whereas the screening for specific genes is more precise.
3. Tokudaia: by Rescuing Y Fragments, Obtain Duality
Another exceptional mammalian group, Ryukyu spiny rats (genus Tokudaia) are endemics of the Nansei Shoto archipelago, Japan. T. osimensis from Amami-Oshima Island and the recently described T. tokunoshimensis from Tokunoshima Island are closely related; both lost a Y chromosome, although their diploid chromosome numbers are different: 2n=25, X0/X0 for T. osimensis and 2n=45, X0/X0 for T. tokunoshimensis [47,48]. The differences (mainly centric fusions) were revealed by comparative chromosome painting [49]. An absence of the Sry gene was proved for T. osimensis [26,50]. However, a small signal of a Y chromosome was revealed in T. osimensis, where a few genes were translocated to the X chromosome (Zfy and Tspy) [51] despite no differences for males and females being revealed by chromosomal painting and by comparative genomic hybridization analyses [52]. These species are endangered, so experiments on T. osimensis cells in vitro look very promising. The results revealed the dual potency for female somatic cells, which differentiated into male germline cells if the male reproductive niche occurred [53]. This experiment highlighted a tremendous significance of supportive cells in sex determination, which was discussed above. T. muenninki (Okinawa spiny rat) retains the Y chromosome unless both sex chromosomes are remarkably large due to fusions with autosomes [54].
4. In Search of a Primary Switch for Mole Voles
Loss of the Sry gene was demonstrated for E. lutescens, E. talpinus and Е. tancrei along with revealing the high homology of the Sry sequence of Е. fuscocapillus to humans and mice [25]. Later, W. Just and colleagues attempted to detect this gene in the genomic DNA of E. lutescens males using Sry probes of distinct species, changing hybridization conditions, etc., but got no specific signal [55]. Recently [56], we re-checked all five species (E. alaicus was unstudied before) and obtained unexpected results. In accordance with previous studies, we did not identify any Sry-similar sequences for males and females of E. lutescens, E. talpinus, Е. tancrei, and, for the first time, E. alaicus. Surprisingly, the presence of a highly conservative HMG box Sry was revealed not only in males but also in females of Е. fuscocapillus. Moreover, in different females, we detected variations in fragments of the HMG box from full (203 bp) to short ones (138 bp), and even an absence of it (in a single female). We suspected that the Sry gene exists in male and female genomes of Е. fuscocapillus in multiple copies as a pseudogene; therefore, its functionality is questionable in this species.
Vogel et al. [57] hypothesized that sex determination in Ellobius species, which lost the Sry gene, might start by mutant alleles of some genes, which usually act downstream of the Sry. For studying the possible role of the genes, segregation analysis was developed. First, fragments of studied genes were amplified by PCR, cloned, and sequenced, and then polymorphic/biallelic markers were searched and screened in at least three generations of families of mole voles of no less than 20 specimens. The same strategy was used in mole voles for the main genes in the sex determination network: SOX9, SF1 (Steroidogenic factor 1 or Nr5a1 nuclear receptor subfamily 5 group A member 1), Sox3 (SRY-box 3), Atrx (alpha thalassemia/mental retardation syndrome X-linked), Nr0b1 (nuclear receptor subfamily 0 group B member 1), Ar (androgen receptor), Foxl2/Pisrt1 (Polled Intersex Syndrome Regulated Transcript 1 (Non-Protein Coding RNA)), and Dmrt1 [25,55,58,59]. No one demonstrated co-segregation of marker alleles with the sex of animals, therefore, a primary sex-determining function was excluded for all mentioned genes in E. lutescens and E. tancrei, i.e., species with X0 or XX sex chromosomes in males and females.
A precise gene expression regulation is essential for sex determination; an illustration is the SOX9 gene, which is involved in numerous processes during development, as well as sex determination. It is known that Sry, together with SF1, binds to the testis-specific enhancer core element (TESCO) of the Sox9 gene to upregulate its expression [60]. Sox9 protein normally starts a genetic cascade in the bipotential somatic precursor cells, which develop into Sertoli cells, resulting in the development of testes. In case of the upregulation of Sox9, the foetal gonads develop as ovaries. Stability of the gene and an enhancer structure are needed because transcription factors should recognize specific binding sites in enhancers for regulation of the gene expression [61]. Nevertheless, in amazing Ellobius and Tokudaia, different deletions were found in the same testis-specific enhancer core element (TESCO) of Sox9. In Tokudaia, a deletion in TESCO was detected in XO species without the Sry gene and in the XY species with multiple Sry gene copies in T. muenninki [62]. In four studied species of mole voles, including Sry-positive species E. fuscocapillus, a 14-bp deletion was detected in the highly conserved module of the TESCO [63]. A deletion in the TESCO might lead to the gene Sox9 upregulation unless a recent paper highlighted more appropriate enhancers for regulating the Sox9 in early development of testis [64]. Nevertheless, the functionality of the sex determination gene network, starting with Sry – SOX9 interactions, seemed doubtful in E. fuscocapillus.
It is possible that E. fuscocapillus restored their sex determination mechanisms by down-regulating the SOX9 expression, as Bagheri-Fam et al. [63] hypothesized. The same method of solving the problem was proposed for Tokudaia species, in which numerous copies of the Cbx2 gene might be involved in male sex determination, providing proper SOX9 regulation [65]. No data for Cbx2 was found in Ellobius yet, and this assumption needs to be checked.
Chandra [66] supposed that sex might be epigenetically determined in E. lutescens with a single X chromosome by transferring an imprinted X from mother to daughter and from father to son. However, analysis of specific microsatellite markers revealed their chaotic inheritance in three generations of mole voles, which was clearly demonstrated to be erroneous of the supposition [55]. The last hope was a whole genome study; a large group of researchers was involved in the project, the data were published and appeared to be rather discouraging [67]. Genomes of only two species, E. lutescens (male and female) and E. talpinus (female) were successfully sequenced and de novo assembled. Several known sex determination genes were recognized in male and female genomes of E. lutescens, and no new candidate sex-determining genes were revealed. To date, the enigma of sex determination in E. lutescens and E. talpinus remains unresolved.
5. The Loss of a Y Chromosome is Not a Deadlock
Although the whole-genome data did not answer the question of sex determining genes, it opened a way to study genes, which are usually located on mammalian Y chromosomes, in Ellobius species lacking this chromosome. Mulugeta et al. [67] detected the genes Zfy1/2 (zinc finger protein Y-linked), Eif2s3y (eukaryotic translation initiation factor 2, subunit 3, structural gene Y-linked), Ssty (spermiogenesis specific transcript on the Y) in E. lutescens and E. talpinus, but Usp9y (ubiquitin specific peptidase 9 Y-linked) was determined in E. lutescens only. FISH demonstrated that the Usp9y and Zfy have been translocated to the X chromosome, so a Y was partly rescued. Although the genes Eif2s3y, Zfy, and Ssty were identified in males and females, the expression of these genes was detected in testes only. We sequenced fragments of the Eif2s3y of all Ellobius species, both males and females, and revealed, that an exonic part of Eif2s3y was demonstrated to identify up to 88% with the gene described for T. muenninki, a species with a Y chromosome [56]. The non-exonic sequenced part was highly variable compared to Ellobius species but was identical in males and females for each species. This part of an Eif2s3y gene (206 bp) in all Ellobius species appeared to be a domesticated fragment of non-coding transposable short interspersed nuclear elements (SINEs) B2–B4 (http://www.repeatmasker.org).
In E. fuscocapillus, Eif2s3y exonic sequences were also identified in males and females. Along with Sry fragments in females, the presence of Eif2s3y proves the suggestion about the translocation of some Y-linked genes to the X chromosome in this species. The detection of the spermatogonial proliferation factor Eif2s3y in male and female genomes is especially interesting because it was shown that it is one of the two essential factors for spermatogenesis genes (the other one is the Sry, see [68]). With high probability, genes lost their primary function, but the data obtained in distinct species confirmed the independent duplications and translocations of some fragments from the ancestral Y to X chromosomes and, possibly, autosomes.
Such a method of compensation for Y degeneration was executed, not only in different Ellobius but also in Tokudaia species.
6. Dosage Compensation in a Case with a Single X Chromosome
The hypothesis of the necessity of dosage compensation for sex chromosomes declares the necessity to balance the gene products in somatic cells of males and females [69,70,71]. It is reliable, at least in part, in mammals with an XX-XY system, although its universal significance is controversial [72].
For Е. lutescens, possessing a single X chromosome, 50% zygotic mortality was detected; zygotes without a sex chromosome, 2n = 16, 00 is lethal, and, unexpectedly, the regular mammals variant with 2n = 18, ХХ is lethal too [73]. The latter variant may be lethal due to insufficient X chromosome inactivation. The system of a sex chromosome dosage compensation is not well-studied to date, and it is suggested that the core event is the initiation of X-chromosome inactivation by nuclear long noncoding RNA (lncRNA) Xist (X inactive specific transcript) [74,75,76].
The structure of the Xist gene was studied in Ellobius and Tokudaia. In Ellobius, the studied fragment of the Xist gene had genus-specific changes, and a deletion, which is not present in Е. fuscocapillus with heteromorphic sex chromosomes [55]. A deletion should lead to the functionality loss of the Xist and might be a source of lethality of embryos with 2n = 18, ХХ in E. lutescens. However, the question remains open on how males and females of three species survive with XX in both sexes.
Numerous mutations were detected in the Xist gene of the Ryukyu spiny rat T. osimensis, which should lead to the loss of a gene function in the XO/XO species. In another species, the Okinawa spiny rat, T. muenninki, with XX/XY and a neo-X obtained by fusion with an autosome, Xist RNAs were expressed in females [77]. Therefore, species with a single X chromosome do not need any mechanisms for X inactivation, their Xist gene remained in the genome, accumulated mutations and was degraded.
7. X0 and XX Sex Chromosomes in Meiosis
Meiosis, a crucial process of development, exhibits significant sexual dimorphism in features of gametogenesis in the two sexes, and the longevity and timing of meiosis differs in females and males. In females, meiotic prophase I starts in germ cells of the ovary during foetal development, undergoes an arrest at the dictyotene stage and ends in adults; the last stages occur after fertilization. In males, meiotic prophase I starts in adult and occurs throughout most of their life, continuously providing gametes.
A diversity of sex chromosomes in Ellobius establishes distinct meiotic patterns. E. fuscocapillus develops a classical system for Eutherians, with a large submetacentric X chromosome and a small acrocentric Y. In the middle pachytene, Y achieved a complete synapsis with X and quickly underwent early complete desynapsis during the late pachytene-early diplotene [42,45,56]. Early desynapsis is uncommon for mammals and might be a sign of the lack of recombination, and further studies are needed. This feature resulted in an unusual ‘typical’ XX-XY system of E. fuscocapillus.
A single submetacentric X chromosome in males and females of E. lutescens is not similar to the X of E. fuscocapillus [42]. the X chromosome of E. lutescens is easy to distinguish as a single univalent in the meiotic prophase I. Using electron microscopy, we revealed that in the pachytene, the Х‐univalent became thicker and developed multiple axes and flexures (‘hairpins’), and their structure was similar to synaptonemal complex. Such structures could be interpreted as possible sites for synapsis, but more evidence is needed for any conclusions. One or two enigmatic round bodies are usually located near the univalent. They are electron-dense, DAPI-positive and H2AFX-negative, and its functions are still unknown [56].
The functional differences were demonstrated for male and female XX chromosomes of E. talpinus and E. tancrei. These acrocentric XX chromosomes with identical G-band morphology underwent a complete synapsis during the pachytene I in females, but in males, XX synapsed and recombined only in the short telomeric regions [56,78,79]. The male XX chromosomes in E. talpinus and E. tancrei formed a typical sex body, which is similar to the XY body in other mammalian males, including E. fuscocapillus. One of the X axes has an electron-dense, SUMO1 (small ubiquitin-related modifier 1)- and DAPI-positive, H2AFX (H2A histone family member X)-negative round structure, which we previously called the dense nucleolar body [80], nucleolus-like body [42,79] or chromatin body [78], respectively. Thus, the two X chromosomes in prophase I are not identical to each other. We suppose that asynapsis between isomorphic XX in males may occur due to asynchronous epigenetic chromatin changes [79]. This functional distinctness in males is presumably an evolutionarily new one, and it might be a start of neo-sex chromosomes in mole voles with two isomorphic X chromosomes.
8. Y Loss in Ellobius: How Many Times?
We hypothesized two independent losses of the Y chromosome for E. lutescens, X0 and E. talpinus-E. tancrei, XX when revealing the specific behaviour of sex chromosomes in meiosis [56,79,81]. Mulugeta et al. [67] argued two independent losses of the Y chromosome for two studied species, E. lutescens and E. talpinus, and proposed a predisposition for the development of a new sex-determination system in the common ancestor of all mole voles. A study of all five species made the scheme more complex [56] (Figure 1).
The 14-bp deletion in TESCO is common for both subgenera of Ellobius; therefore, we started the reconstruction of their evolutionary history from this event (Figure 1). Unless the role of this enhancer in Sox9 regulation during early gonadogenesis was overestimated [64], changes in the enhancer structure could be an accidentally discovered consequence of the genome rearrangement, caused by an unknown crucial event. We assumed that after those genomic disturbances, species evolved in different ways due to their genome specificity. We suppose an independent loss of Y chromosomes in E. lutescens and in the subgenus Ellobius after the separation of two subgenera (Figure 1).
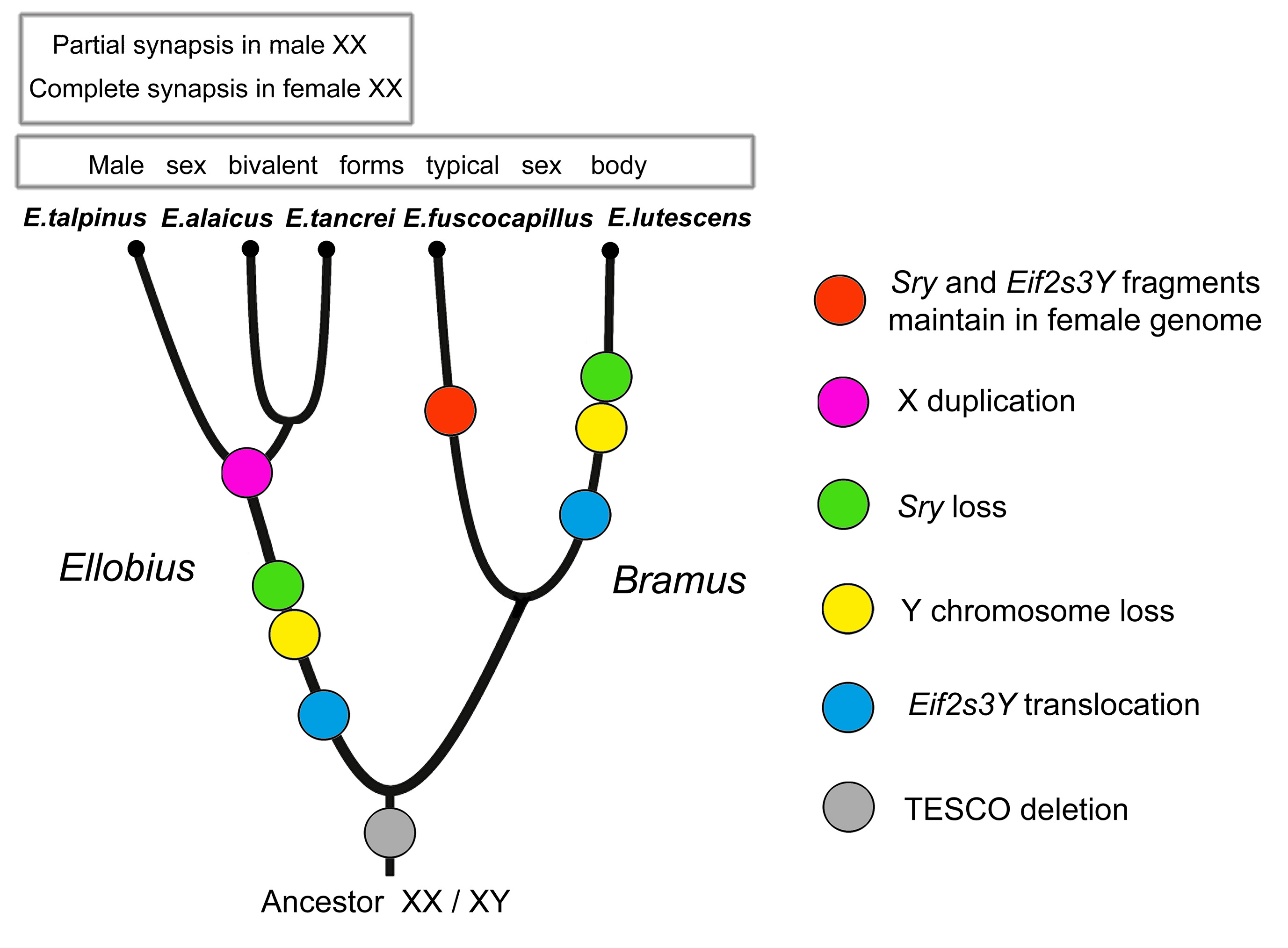
Figure 1 Evolution of sex chromosomes in mole voles Ellobius. Phylogenetic reconstruction is based on data of sex chromosomes and genes involved in sex determination and spermatogenesis.
Moreover, according to different changes in genes, species of two subgenera (Bramus and Ellobius) underwent distinct events of copying Y-linked genes and the loss of the entire Y chromosome. The presence of the Sry fragments in females of E. fuscocapillus, and the difference in structure of Eif2s3y between this species and E. lutescens revealed a high level of divergency in the subgenus Bramus. The comparison of structures of sequenced fragments of Eif2s3y and Eif2s3x genes alongside meiotic features led us to the conclusion that the X chromosome in E. talpinus, E. tancrei and E. alaicus was doubled after translocations of the Y-located genes to the X. Later, XX underwent a new heteromorphization, became functionally different in the central region, and restricted synapsis to telomeric areas in males (Figure 1).
Evolution of the X chromosomes in mole voles is different in subgenera too. In E. fuscocapillus and E. lutescens, the Xs are submetacentrics are different in size and G-band morphology. In subgenera Ellobius, the X chromosomes are large acrocentrics, undistinguishable by G-band in different species [42], and these chromosomes demonstrate functional distinctness in male and female meiosis [78,79].
If a Y chromosome disappeared in some mole voles, might a single X in E. lutescens be lost too? This idea was discussed by W. Just (personal communication). He supposed a degeneration of the single X chromosome, which lost a partner for recombination, because a loss of recombination may lead to the accumulation of mutations. If mutations appeared to be deleterious, a mechanism of "Muller's ratchet" should eliminate species. However, even if non-deleterious mutations accumulate, they may inactivate X-linked genes and lead to extinction of the species.
The degeneration of the X chromosome might also be a start for the new cycle of sex chromosome evolution, when neo-sex chromosomes develop from autosomes. In Ellobius with two X chromosomes, we probably observe neo-sex chromosomes, which are morphologically identical, but functionally different.
9. Altered Autosomes and Meiotic Drive
Evolution of both sex chromosomes and autosomes may accompany or initiate speciation [82,83]. In the case of sex chromosomes, as we tried to demonstrate above, the main point for a divergence is an emergence of different patterns in a meiotic behaviour of sex chromosomes. The same is true for autosomes. Inversions could lead to sterility; therefore, such chromosome changes are generally accepted as possible speciation mechanisms [82]. Mice are one of the most well-studied and fruitful groups for studying speciation by chromosome changes [84]. Many different fusions lead to different sets of metacentrics in karyotypes, along with different a combination of fused chromosomes. The same situation was found in Ellobius tancrei, one of the mole voles species with XX chromosomes in males and females [80,85,86,87,88,89]. The most intriguing question is how numerous chromosomal changes could originate? A chain process, i.e., consequential fusions of acrocentrics, seems to be the simplest solution [90], but in the case of partial fusions, monobrachial homology is not an appropriate answer. The most promising is the idea that chromosome changes occur during gametogenesis; in such a case, we should conclude that the main mechanism of their origin must relate to meiosis. This hypothesis is rather problematic to prove because appropriate models and concepts of fixing changes should be developed. Robertsonian translocations were noticed as evolutionary neutral for a long time. Such a view may be true in a case of translocations, when hybrids are heterozygous by one translocation. In such a ‘simple’ case, a trivalent emerges in the pachytene stage, the meiosis ends with two types of gametes: with single metacentric or two acrocentrics. In the case of selective advantage for organisms with Rb translocation, it has a chance to spread; otherwise, it will keep as an occasional variant. One of the leading hypotheses of the preferential transmission of chromosomes with strong centromeres against chromosomes with weak centromeres [91,92] appears inappropriate because the molecular basis for centromere identity is elusive. These parts of the chromosomes are not supported by specific DNA, and are presumably epigenetically regulated [93,94]. It is still unknown why and how such a rigid structure as a centromere might have originated through obscure epigenetic mechanisms and survive under natural selection in evolution.
An even more complicated scenario is possible for cases with numerous translocations, especially when distinct translocations emerge in different populations, or when the translocations appeared to be partly homological. The homology by a single branch was named ‘monobrachial’ [95]. There are numerous cases for such translocations in Mus [96], Rhogeessa tumida [97], Sorex araneus [98], Rattus sordidius [99], rock wallabies [100] and others. In bats, monobrachial translocations were possibly mainstream for speciation [101]. Why did emergences of partial homology in chromosome fusions lead to divergence? The meiosis appeared to be a main barrier for spreading the re-arranged chromosomal sets. In hybrids, during prophase I, the homological chromosomes or their parts synapse in the case of monobrachial homology, and different tetra-, pentavalents or even more complicated figures and chains are puzzled. The proper segregation for the complexes is rather difficult and often leads to abnormal chromosomal sets in metaphase II. The meiotic drive as a biased transmission of genetic variants might be a result of meiotic events during gametogenesis. In female meiosis, homologous chromosomes might be differentially transmitted to the egg or polar bodies. In males, a specific genetic element might disturb the function of sperm that reduces fertility [102,103,104].
10. Meiotic Troubleshooting in Hybrids with Partial Chromosome Homology
Chromosomal rearrangements, including Robertsonian translocations, can lead to the formation of new balanced karyotypes, nevertheless changing the architectonics of the nucleus. A concept of chromosome territories proposes a non-random distribution of chromosomes in nuclei; a nuclear architecture constitutes the basis for gene expression regulation [105]. The main output is that a transformation of chromatin structures may alter the genetic system of the species due to the modulation accessibility of transcription factors to DNA binding sites, thus regulating gene expression. Robertsonian fusions restructure the organization of the nuclei, especially in the case when chromosome territories of fused chromosomes are located far from each other. The differences in locations may promote the fusions and encourage the meiotic drive in favour of changed chromosomes [106,107] or against them [108].
In diverse model crossings, which were made for Mus domesticus, distinct meiotic disturbances were revealed. For example, different multivalents formed associations with a sex bivalent in prophase I [109,110]. To evaluate the possible input of the translocations to species diversification, we studied several cases of monobrachial homology in E. tancrei, a species with numerous chromosomal forms. The homology of chromosomes was verified by chromosome painting, because G-banding in some cases was not precise enough. The scheme of our experimental hybridization is demonstrated in Figure 2.
We crossed a form with 2n = 50, two pairs of Rbs: 2Rb(2.18) and 2Rb(5.9), which was nicknamed 'Voidara' according to the closest settlement in the Surkhob River Valley (Figure 2b), with two different forms: another form with 2n = 50, distinct two pairs of Rbs: 2Rb(4.12) and 2Rb(9.13), named ‘Khodza Obi-Garm’ (Figure 2a), and 2n = 48, three pairs of metacentrics: 2Rb(2.11), 2Rb(5.9), and 2Rb(3.18) (Figure 2c). F1 hybrids of the first crossing had 2n=50 and four distinct Rbs (1Rb(2.18), 1Rb(4.12), 1Rb(5.9), and 1Rb(9.13), two of them with monobrachial homology (Figure 2d). In meiotic prophase I, two trivalents and a tetravalent were revealed (Figure 2f). The second crossing resulted in F1 hybrids with 2n=49 and five Rb metacentrics (2Rb(5.9), 1Rb(2.11), 1Rb(2.18) and 1Rb(3.18)), where three of them obtained partial or monobrachial homology (Figure 2e); therefore, a chain (pentavalent) was detected in meiotic prophase I (Figure 2g). These chromosomal forms do not interbreed in nature because they inhabit geographically separated areas in the Pamir-Alay Mountains. In the laboratory, no behavioural differences or preferences for the forms were revealed. Hybrids of the first generation had lower fertility but did not exhibit any health problems, and their longevity was the same as parental ones (up to seven years). Hybrid fertility increased starting in the third generation after the deep inbreeding depression in the first generation [111,112].
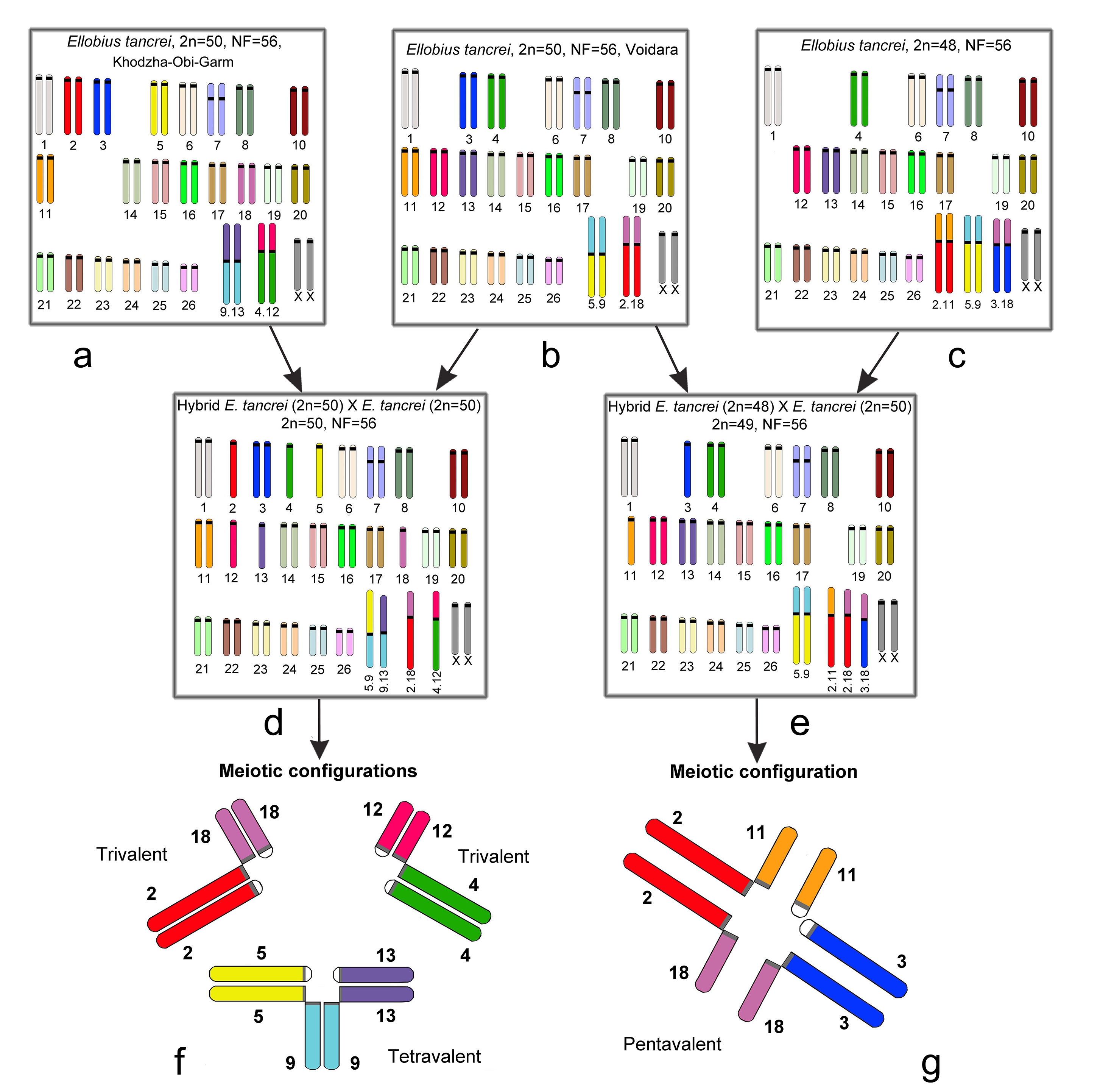
Figure 2 Scheme of experimental hybridization in E. tancrei. a. form ‘Khodza Obi-Garm’, 2n = 50, 2Rb(4.12) and 2Rb(9.13); b. form 'Voidara', 2n = 50, 2Rb(2.18), and 2Rb(5.9); c. form 2n = 48, 2Rb(2.11), 2Rb(5.9), and 2Rb(3.18); d. F1 hybrid with 2n=50, 1Rb(2.18), 1Rb(4.12), 1Rb(5.9), and 1Rb(9.13); e. F1 hybrid with 2n=49, 2Rb(5.9), 1Rb(2.11), 1Rb(2.18) and 1Rb(3.18); f. the meiotic prophase I of F1 hybrid (2d), with two trivalents and a tetravalent present; g. the meiotic prophase I of the F1 hybrid (2e), and the pentavalent consists of 3 Rb metacentrics with monobrachial homology and 2 acrocentrics.
A particularly interesting output was a meiotic solution for heterozygotes by Rb metacentrics. The most common disturbance was delayed synapses, which resumed later if compared to the homologous crossings. The late synaptic adjustments nevertheless provide a proper segregation of chromosomes and normal sets in the gametes. The question of a lower rate of recombination due to delayed synapses is still open; apparently, it may lead to negative consequences in an evolutionary perspective. The role of monobrachial fusions in speciation presumably depends on karyotype organization, and in some cases, may lead to fast divergence, even in several generations, and especially in the case of the physical isolation of hybrids from the parental forms.
11. Conclusion
Loss of the Sry gene and hypothetical alteration of the regulation of its target SOX9 gene in several Ellobius and Tokudaia species raises a question of universality in the sex determining gene network in placental mammals. A plausible way is an origin of compensatory mutations to correct the disrupted testis determining pathway. One of the possible candidates is the Cbx2 gene, which might be involved in male sex determination, providing a proper SOX9 regulation in Tokudaia [65]. Different genes were studied as a potential primary sex factor in Ellobius and Tokudaia, but to date, we do not know how the early development of gonads occurs. Distinctness in the meiotic behaviour of sex chromosomes in Ellobius species with two X chromosomes (E. talpinus and E. tancrei) may be evaluated as a first sign of emergence of new sex chromosomes when X chromosomes in females are fully homological, but in males, these chromosomes demonstrate as typical for heteromorphic sex chromosome behaviour. In a case of sex chromosome origin, and in inheritance of changed autosomes, meiosis appeared to be an essential process for emergence, altering the inheritance of newly originated sex and autosomes [113].
An enigma of when and how Robertsonian translocations may originate will probably get an answer soon. Now, it became possible to study molecular mechanisms of genome instability, and first data on the emergence of translocations in germ lines appeared (see review in [114]). The changes rise in meiosis due to double-strand breaks, and meiosis is also involved in next evolutionary steps through meiotic drive mechanism.
Mole voles Ellobius, as exceptions to the rule, are a fruitful model for further studying the evolution of sex determination and chromosomal speciation.
Author Contributions
Authors contributed equally to this work.
Funding Source
This study was supported in part by the Russian Foundation for Basic Researches N 17-04-00618 and N 15-29-02649.
Competing Interests
The authors have declared that no competing interests exist.
References
- Wilhelm D, Palmer S, Koopman P. Sex determination and gonadal development in mammals. Physiol Rev. 2007; 87: 1-28. [CrossRef]
- Bachtrog D, Mank JE, Peichel CL, Kirkpatrick M, Otto SP, Ashman TL, et al. Sex determination: why so many ways of doing it? PLoS Biol. 2014; 12: e1001899. [CrossRef]
- Capel B. Vertebrate sex determination: evolutionary plasticity of a fundamental switch. Nat Rev Genet. 2017; 18: 675-689. [CrossRef]
- Rens W, Grützner F, O'Brien PC, Fairclough H, Graves JA, Ferguson-Smith MA. Resolution and evolution of the duck-billed platypus karyotype with an X1Y1X2Y2X3Y3X4Y4X5Y5 male sex chromosome constitution. P Natl Acad Sci USA. 2004; 101: 16257-16261. [CrossRef]
- Rens W, O'Brien PC, Grützner F, Clarke O, Graphodatskaya D, Tsend-Ayush E, et al. The multiple sex chromosomes of platypus and echidna are not completely identical and several share homology with the avian Z. Genome Biol. 2007; 8: R243. [CrossRef]
- Bininda-Emonds OR, Cardillo M, Jones KE, MacPhee RD, Beck RM, Grenyer R, et al. The delayed rise of present-day mammals. Nature. 2007; 446: 507-512. [CrossRef]
- Cortez D, Marin R, Toledo-Flores D, Froidevaux L, Liechti A, Waters PD, et al. Origins and functional evolution of Y chromosomes across mammals. Nature. 2014; 508: 488-493. [CrossRef]
- Ayling LJ, Griffin DK. The evolution of sex chromosomes. Cytogenet Genome Res. 2002; 99: 125-140. [CrossRef]
- Charlesworth D, Charlesworth B, Marais G. Steps in the evolution of heteromorphic sex chromosomes. Heredity. 2005; 95: 118-128. [CrossRef]
- Bellott DW, Hughes JF, Skaletsky H, Brown LG, Pyntikova T, Cho TJ, et al. Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators. Nature. 2014; 508: 494-499. [CrossRef]
- McElreavey K, Vilain E, Cotinot C, Payen E, Fellous M. Control of sex determination in animals. FEBS J. 1993; 218: 769-783. [CrossRef]
- Gubbay J, Collignon J, Koopman P, Capel B, Economou A, Münsterberg A, et al. A gene mapping to the sex-determining region of the mouse Y chromosome is a member of a novel family of embryonically expressed genes. Nature. 1990; 346: 245-250. [CrossRef]
- Sinclair AH, Berta P, Palmer MS, Hawkins JR, Griffiths BL, Smith MJ, et al. A gene from the human sex-determining region encodes a protein with homology to a conserved DNA-binding motif. Nature. 1990; 346: 240-244. [CrossRef]
- Koopman P, Gubbay J, Vivian N, Goodfellow P, Lovell-Badge R. Male development of chromosomally female mice transgenic for Sry. Nature. 1991; 351: 117-121. [CrossRef]
- Nishino K, Hattori N, Tanaka S, Shiota K. DNA methylation-mediated control of Sry gene expression in mouse gonadal development. J Biol Chem. 2004; 279: 22306-22313. [CrossRef]
- Kuroki S, Matoba S, Akiyoshi M, Matsumura Y, Miyachi H, Mise N, et al. Epigenetic regulation of mouse sex determination by the histone demethylase Jmjd1a. Science. 2013; 341: 1106-1109. [CrossRef]
- Garcia-Moreno SA, Plebanek MP, Capel B. Epigenetic regulation of male fate commitment from an initially bipotential system. Mol Cell Endocrinol. 2018. 468: 19-30. [CrossRef]
- Parma P, Radi O, Vidal V, Chaboissier MC, Dellambra E, Valentini S, et al. R-spondin1 is essential in sex determination, skin differentiation and malignancy. Nat Genet. 2006; 38: 1304–1309. [CrossRef]
- Biason-Lauber A, Chaboissier MC. Ovarian development and disease: the known and the unexpected. Semin cell Dev Biol. 2015; 45: 59-67. [CrossRef]
- Uhlenhaut NH, Jakob S, Anlag K, Eisenberger T, Sekido R, Kress J, et al. Somatic sex reprogramming of adult ovaries to testes by FOXL2 ablation. Cell. 2009; 139: 1130-1142. [CrossRef]
- Matson CK, Murphy MW, Sarver AL, Griswold MD, Bardwell VJ, Zarkower D. DMRT1 prevents female reprogramming in the postnatal mammalian testis. Nature. 201; 476: 101-104. [CrossRef]
- Barrionuevo FJ, Hurtado A, Kim GJ, Real FM, Bakkali M, Kopp JL, et al. Sox9 and Sox8 protect the adult testis from male-to-female genetic reprogramming and complete degeneration. Elife. 2016; 5: e15635. doi: 10.7554/eLife.15635. [CrossRef]
- Waters PD, Ruiz-Herrera A, Dobigny G, Caldès MG, Robinson TJ. Sex chromosomes of basal placental mammals. Chromosoma. 2007; 116: 511-518. [CrossRef]
- Katsura Y, Kondo HX, Ryan J, Harley V, Satta Y. The evolutionary process of mammalian sex determination genes focusing on marsupial SRYs. BMC Evol Biol. 2018; 18: 3. [CrossRef]
- Just W, Rau W, Vogel W, Akhverdian M, Fredga K, Graves JA, et al. Absence of Sry in species of the vole Ellobius. Nat Genet. 1995; 11: 117-118. [CrossRef]
- Soullier S, Hanni C, Catzeflis F, Berta P, Laudet V. Male sex determination in the spiny rat Tokudaia osimensis (Rodentia: Muridae) is not Sry dependent. Mamm Genome. 1998; 9: 590-592. [CrossRef]
- O'Neill MJ, O'Neill RJ. Sex chromosome repeats tip the balance towards speciation. Mol Ecol. 2018, in press. doi:10.1111/mec.14577 [CrossRef]
- Graves JA. Did sex chromosome turnover promote divergence of the major mammal groups? Bioessays. 2016; 38: 734-743. [CrossRef]
- Pallas PS. Spicilegia zoologica, quibus novae imprimus et obscurae animalium species iconibus, descriptionibus atque commentariis illustrantur cura P.S. Pallas. 1767-1780. Berolini, prostant apud Gottl. August. Langed, 14 fasc in 2 volumes.
- Fischer G. Zoognosia tabulis synopticis illustrata. 1814. Mosquae [Moscow]: Typis Nicolai S. Vsevolozsky, V. 3.
- Musser GG, Carleton MD: Subfamily Arvicolinae, in Wilson DE, Reeder DM (eds): Mammal Species of the World: A Taxonomic and Geographic Reference, ed 3. 2005. Baltimore: Johns Hopkins Univ Press. 956–1039.
- Matthey R. La formule chromosomique et le problème de la détermination sexuelle chez Ellobius lutescens (Rodentia-Muridae-Microtinae). Arch. Julius Klaus-Stift Vererb. Forsch. 1953; 28: 65–73.
- Lyapunova EA, Vorontsov NN. Genetics of Ellobius (Rodentia). I. Karyological characteristics of four Ellobius species. Genetika. 1978; 14: 2012–2024. (In Russian).
- Ivanov VG: Chromosomal complex of common mole vole. Tsitologiia. 1967; 9: 879–883
- Vorontsov NN, Radjabli SI. Chromosomal sets and cytogenetic differentiation of two forms within the superspecies Ellobius talpinus Pall. Tsitologiia. 1967; 9: 848–852. (In Russian).
- Vorontsov NN, Lyapunova EA, Zakarjan GG, Ivanov VG. The karyology and taxonomy of the genus Ellobius (Microtinae, Rodentia), in Vorontsov NN (ed): The Mammals: Evolution, Karyology, Faunistics, Systematics. 1969. Novosibirsk: Nauka. 127–129. (In Russian).
- Romanenko SA, Sitnikova NA, Serdukova NA, Perelman PL, Rubtsova NV, et al: Chromosomal evolution of Arvicolinae (Cricetidae, Rodentia): II. The genome homology of two mole voles (genus Ellobius), the field vole and golden hamster revealed by comparative chromosome painting. Chromosome Res. 2007; 15: 891–897. [CrossRef]
- Bakloushinskaya IY, Matveevsky SN, Romanenko SA, Serdukova NA, Kolomiets OL, Spangenberg VE, et al. A comparative analysis of the mole vole sibling species Ellobius tancrei and E. talpinus (Cricetidae, Rodentia) through chromosome painting and examination of synaptonemal complex structures in hybrids. Cytogenet Genome Res. 2012; 136: 199-207. [CrossRef]
- Romanenko SA, Lemskaya NA, Trifonov VA, Serdyukova NA, O’Brien PC, Bulatova NS, et al. Genome-wide comparative chromosome maps of Arvicola amphibius, Dicrostonyx torquatus, and Myodes rutilus. Chromosome Res. 2016; 24: 145-59. [CrossRef]
- White MJD: An interpretation of the unique sex chromosome mechanism of the rodent Ellobius lutescens. Proc Zool Soc Calcutta, Moorkerjee Mem Vol. 1957; 113–114.
- Matthey R. Un nouveau type de determination chromosomique du sex chez les mammiferes Ellobius lutescens Th. etMicrotus (Chilotus) Bachm (Murides-Microtines). Cell Mol Life Sci. 1958; 14: 240–241. [CrossRef]
- Castro-Sierra E, Wolf U. Studies on the male meiosis of Ellobius lutescens Th. Cytogenet Genome Res. 1968; 7: 241-248. [CrossRef]
- Castro-Sierra E, Wolf U. Replication patterns of the unpaired chromosome No. 9 of the rodent Ellobius lutescens Th. Cytogenet Genome Res. 1967; 6: 268-275. [CrossRef]
- Vogel W, Steinbach P, Djalali M, Mehnert K, Ali S, Epplen JT. Chromosome 9 of Ellobius lutescens is the X chromosome. Chromosoma. 1988; 96: 112-118. [CrossRef]
- Kolomiets OL, Vorontsov NN, Lyapunova EA, Mazurova TF. Ultrastructure, meiotic behavior, and evolution of sex chromosomes of the genus Ellobius. Genetica. 1991; 84:179–189. [CrossRef]
- Just W, Baumstark A, Hameister H, Schreiner B, Reisert I, Hakhverdyan M, et al. The sex determination in Ellobius lutescens remains bizarre. Cytogenet Genome Res. 2002; 96: 146-153. [CrossRef]
- Honda T, Suzuki H, Itoh M. An unusual sex chromosome constitution found in the Amami spinous country-rat, Tokudaia osimensis osimensis. Jpn J Genet. 1977; 52: 247-249. [CrossRef]
- Honda T, Suzuki H, Itoh M, Hayashi K. Karyotypical differences of the Amami spinous country-rats, Tokudaia osimensis osimensis obtained from two neighbouring islands. Jpn J Genet. 1978; 53: 297-299. [CrossRef]
- Nakamura T, Kuroiwa A, Nishida-Umehara C, Matsubara K,Yamada F, Matsuda Y (2007) Comparative chromosome painting map between two Ryukyu spiny rat species, Tokudaia osimensis and Tokudaia tokunoshimensis (Muridae, Rodentia). Chromosome Res 15:799–806 [CrossRef]
- Sutou S, Mitsui Y, Tsuchiya K. Sex determination without the Y chromosome in two Japanese rodents Tokudaia osimensis osimensis and Tokudaia osimensis spp. Mamm Genome. 2001; 12: 17-21. [CrossRef]
- Arakawa Y, Nishida-Umehara C, Matsuda Y, Sutou S, Suzuki H. X-chromosomal localization of mammalian Y-linked genes in two XO species of the Ryukyu spiny rat. Cytogenet Genome Res. 2002; 99: 303-309. [CrossRef]
- Kobayashi T, Yamada F, Hashimoto T, Abe S, Matsuda Y, Kuroiwa A. Exceptional minute sex-specific region in the X0 mammal, Ryukyu spiny rat. Chromosome Res. 2007; 15: 175-187. [CrossRef]
- Honda A, Choijookhuu N, Izu H, Kawano Y, Inokuchi M, Honsho K, et al. Flexible adaptation of male germ cells from female iPSCs of endangered Tokudaia osimensis. Sci Adv. 2017; 3: e1602179 [CrossRef]
- Murata C, Kuroki Y, Imoto I, Kuroiwa A. Ancestral Y-linked genes were maintained by translocation to the X and Y chromosomes fused to an autosomal pair in the Okinawa spiny rat Tokudaia muenninki. Chromosome Res. 2016; 24: 407-419. [CrossRef]
- Just W, Baumstark A, Süss A, Graphodatsky A, Rens W, Schäfer N, et al. Ellobius lutescens: sex determination and sex chromosome. Sex Dev. 2007; 1: 211-221. [CrossRef]
- Matveevsky S, Kolomiets O, Bogdanov A, Hakhverdyan M, Bakloushinskaya I. Chromosomal evolution in mole voles Ellobius (Cricetidae, Rodentia): bizarre sex chromosomes, variable autosomes and meiosis. Genes. 2017; 8: 306. [CrossRef]
- Vogel W, Jainta S, Rau W, Geerkens C, Baumstark A, Correa-Cerro LS, et al. Sex determination in Ellobius lutescens: the story of an enigma. Cytogenet Genome Res. 1998; 80: 214-221. [CrossRef]
- Baumstark A, Akhverdyan M, Schulze A, Reisert I, Vogel W, et al. Exclusion of SOX9 as the testis determining factor in Ellobius lutescens: evidence for another testis determining gene besides SRY and SOX9. Mol Genet Metab. 2001; 72: 61–66. [CrossRef]
- Baumstark A, Hameister H, Hakhverdyan M, Bakloushinskaya I, Just W. Characterization of Pisrt1/Foxl2 in Ellobius lutescens and exclusion as sex-determining genes. Mamm Genome. 2005; 16: 281-289. [CrossRef]
- Sekido R, Lovell-Badge R. Sex determination involves synergistic action of SRY and SF1 on a specific Sox9 enhancer. Nature. 2008; 453: 930-934. [CrossRef]
- Grossman SR, Zhang X, Wang L, Engreitz J, Melnikov A, Rogov P, et al. Systematic dissection of genomic features determining transcription factor binding and enhancer function. P Natl Acad Sci. 2017; 114: E1291–E1300. [CrossRef]
- Kimura, R.; Murata, C.; Kuroki, Y.; Kuroiwa, A. Mutations in the testis-specific enhancer of SOX9 in the SRY independent sex-determining mechanism in the genus Tokudaia. PLoS ONE. 2014; 9: e108779. [CrossRef]
- Bagheri-Fam S, Sreenivasan R, Bernard P, Knower KC, Sekido R, Lovell-Badge R, et al. Sox9 gene regulation and the loss of the XY/XX sex-determining mechanism in the mole vole Ellobius lutescens. Chromosome Res. 2012; 20: 191-199. [CrossRef]
- Gonen N, Futtner CR, Wood S, Garcia-Moreno SA, Salamone IM, Samson SC, et al. Sex reversal following deletion of a single distal enhancer of Sox9. Science. 2018; 360: 1469-1473. [CrossRef]
- Kuroiwa A, Handa S, Nishiyama C, Chiba E, Yamada F, Abe S, et al. Additional copies of CBX2 in the genomes of males of mammals lacking SRY, the Amami spiny rat (Tokudaia osimensis) and the Tokunoshima spiny rat (Tokudaia tokunoshimensis). Chromosome Res 2011; 19: 635-644. [CrossRef]
- Chandra SH. Another way of looking at the enigma of sex determination in Ellobius lutescens. Curr Sci. 1999; 76: 1072-1073.
- Mulugeta E, Wassenaar E, Sleddens-Linkels E, van IJcken WF, Heard E, Grootegoed JA, et al. Genomes of Ellobius species provide insight into the evolutionary dynamics of mammalian sex chromosomes. Genome Res. 2016; 26: 1202-1210. [CrossRef]
- Yamauchi Y, Riel JM, Ruthig VA, Ortega EA, Mitchell MJ, Ward MA. Two genes substitute for the mouse Y chromosome for spermatogenesis and reproduction. Science. 2016; 351: 514-516. [CrossRef]
- Lyon MF. Gene action in the X-chromosome of the mouse (Mus musculus L.) Nature 1961; 190: 372–373. [CrossRef]
- Lyon MF. X-chromosome inactivation: a repeat hypothesis. Cytogenet Cell Genet. 1998; 80: 133–137. [CrossRef]
- Ohno, S. Sex chromosomes and sex-linked genes. 1967. Springer-Verlag. [CrossRef]
- Gu L, Walters JR. Evolution of sex chromosome dosage compensation in animals: a beautiful theory, undermined by facts and bedeviled by details. Genome Biol Evol. 2017; 9: 2461-2476. [CrossRef]
- Lyapunova EA, Vorontsov NN, Zakarjan GG. Zygotic mortality in Ellobius lutescens (Rodentia: Microtinae). Cell Mol Life Sci (Experientia). 1975; 31: 417-418. [CrossRef]
- Penny GD, Kay GF, Sheardown SA, Rastan S, Brockdorff N. Requirement for Xist in X chromosome inactivation. Nature. 1996; 379: 131-137. [CrossRef]
- Okamoto I, Patrat C, Thépot D, Peynot N, Fauque P, Daniel N, et al. Eutherian mammals use diverse strategies to initiate X-chromosome inactivation during development. Nature. 2011; 472: 370-374. [CrossRef]
- Wutz A. Gene silencing in X-chromosome inactivation: advances in understanding facultative heterochromatin formation. Nat Rev Genet. 2011; 12: 542-553. [CrossRef]
- Zushi H, Murata C, Mizushima S, Nishida C, Kuroiwa A. Unique XCI evolution in Tokudaia: initial XCI of the neo-X chromosome in Tokudaia muenninki and function loss of XIST in Tokudaia osimensis. Chromosoma. 2017; 126: 741-751. [CrossRef]
- Kolomiets OL, Matveevsky SN, Bakloushinskaya IY. Sexual dimorphism in prophase I of meiosis in the Northern mole vole (Ellobius talpinus Pallas, 1770) with isomorphic (XX) chromosomes in males and females. Comp Cytogenet. 2010; 4: 55–66. [CrossRef]
- Matveevsky S, Bakloushinskaya I, Kolomiets O. Unique sex chromosome systems in Ellobius: How do male XX chromosomes recombine and undergo pachytene chromatin inactivation? Sci Rep-UK. 2016; 6: 29949. [CrossRef]
- Bogdanov YF, Kolomiets OL, Lyapunova EA, Yanina IY, Mazurova TF. Synaptonemal complexes and chromosome chains in the rodent Ellobius talpinus heterozygous for ten Robertsonian translocations. Chromosoma. 1986; 94: 94-102. [CrossRef]
- Matveevsky SN. Signs of sexual dimorphism in meiosis and karyotype variability of mole vole Ellobius (Rodentia, Mammalia). Ph.D. Thesis, NI Vavilov Institute of General Genetics of Russian Academy of Science, Moscow, 2011: 1–172.(in Russian)
- King M. (1993) Species evolution. The role of chromosome change. Cambridge: Cambridge University Press. 336 pp.
- Dobigny G, Britton‐Davidian J, Robinson TJ. Chromosomal polymorphism in mammals: an evolutionary perspective. Biol Rev. 2017; 92: 1-21. [CrossRef]
- Castiglia R. Sympatric sister species in rodents are more chromosomally differentiated than allopatric ones: implications for the role of chromosomal rearrangements in speciation. Mammal Rev. 2014; 44: 1-4. [CrossRef]
- Lyapunova EA, Vorontsov NN, Korobitsyna KV, Ivanitskaya EY, Borisov YM, Yakimenko LV, et al. A Robertsonian fan in Ellobius talpinus. Genetica. 1984; 52: 239-247. [CrossRef]
- Lyapunova EA, Ivnitskii SB, Korablev VP. Yanina IYu. Complete Robertsonian fan of the chromosomal forms in the mole-vole superspecies Ellobius talpinus. Doklady Akademii Nauk SSSR. 1984; 274: 1209-1213. (In Russian).
- Lyapunova EA, Bakloushinskaya IY, Saidov AS, Saidov KK. Dynamics of chromosome variation in mole voles Ellobius tancrei (Mammalia, Rodentia) in Pamiro-Alay in the period from 1982 to 2008. Russ J Genet. 2010; 46: 566-571. [CrossRef]
- Bakloushinskaya IY, Romanenko SA, Graphodatsky AS, Matveevsky SN, Lyapunova EA, Kolomiets OL. The role of chromosome rearrangements in the evolution of mole voles of the genus Ellobius (Rodentia, Mammalia). Russ J Genet. 2010; 46: 1143-1145. [CrossRef]
- Bakloushinskaya I, Romanenko SA, Serdukova NA, Graphodatsky AS, Lyapunova EA. A new form of the mole vole Ellobius tancrei Blasius, 1884 (Mammalia, Rodentia) with the lowest chromosome number. Comp Cytogenet. 2013; 7: 163-169. [CrossRef]
- White MJD. Chain processes in chromosomal speciation. Syst Zool. 1978; 27: 285-298. [CrossRef]
- Henikoff S, Malik HS. Centromeres: selfish drivers. Nature. 2002; 417: 227. [CrossRef]
- Chmátal L, Gabriel SI, Mitsainas GP, Martínez-Vargas J, Ventura J, Searle JB, et al. Centromere strength provides the cell biological basis for meiotic drive and karyotype evolution in mice. Curr Biol. 2014; 24: 2295-2300. [CrossRef]
- McKinley KL, Cheeseman IM. The molecular basis for centromere identity and function. Nat Rev Mol Cell Biol. 2016; 17: 16-29. [CrossRef]
- Rosin LF, Mellone BG. Centromeres drive a hard bargain. Trends Genet. 2017; 33: 101-117. [CrossRef]
- Baker RJ, Bickham JW. Speciation by monobrachial centric fusions. P Natl Acad Sci. 1986; 83: 8245-8248. [CrossRef]
- Nunes AC, Catalan J, Lopez J, da Graça Ramalhinho M, da Luz Mathias M, Britton-Davidian J. Fertility assessment in hybrids between monobrachially homologous Rb races of the house mouse from the island of Madeira: implications for modes of chromosomal evolution. Heredity. 2011; 106: 348-356. [CrossRef]
- Baker RJ, Bickham JW, Arnold ML. Chromosomal evolution in Rhogeessa (Chiroptera: Vespertilionidae): possible speciation by centric fusions. Evolution. 1985; 39: 233-243. [CrossRef]
- White TA, Bordewich M, Searle JB. A network approach to study karyotypic evolution: the chromosomal races of the common shrew (Sorex araneus) and house mouse (Mus musculus) as model systems. Syst Biol. 2010; 59: 262-276. [CrossRef]
- Baverstock PR, Gelder M, Jahnke A. Chromosome evolution in Australian Rattus—G-banding and hybrid meiosis. Genetica. 1983; 60: 93-103. [CrossRef]
- Potter S, Bragg JG, Blom MP, Deakin JE, Kirkpatrick M, Eldridge MD, et al. Chromosomal speciation in the genomics era: Disentangling phylogenetic evolution of rock-wallabies. Front Genet. 2017; 8: 10. [CrossRef]
- Sotero-Caio CG, Baker RJ, Volleth M. Chromosomal evolution in Chiroptera. Genes. 2017; 8: 272. [CrossRef]
- Sandler L, Novitski E. Meiotic drive as an evolutionary force. Am Nat. 1957; 91: 105-110. [CrossRef]
- de Villena FP, Sapienza C. Female meiosis drives karyotypic evolution in mammals. Genetics. 2001; 159: 1179-1189.
- Lindholm AK, Dyer KA, Firman RC, Fishman L, Forstmeier W, Holman L, et al. The ecology and evolutionary dynamics of meiotic drive. Trends Ecol Evol. 2016; 31: 315-326. [CrossRef]
- Cremer T, Cremer M. Chromosome territories. CSH Perpect Biol. 2010; 2: a003889. [CrossRef]
- Parada LA, Misteli T. Chromosome positioning in the interphase nucleus. Trends Cell Biol. 2002; 12: 425-432. [CrossRef]
- Malik HS, Henikoff S. Conflict begets complexity: the evolution of centromeres. Curr Opin Genet Dev. 2002; 12: 711-718. [CrossRef]
- Nachman MW, Searle JB. Why is the house mouse karyotype so variable? Trends Ecol Evol. 1995; 10: 397-402. [CrossRef]
- Berríos S, Fernández-Donoso R, Ayarza E. Synaptic configuration of quadrivalents and their association with the XY bivalent in spermatocytes of Robertsonian heterozygotes of Mus domesticus. Biol Res. 2017; 50: 38. [CrossRef]
- Berríos S, Fernández-Donoso R, Page J, Ayarza E, Capanna E, Solano E, et al. Hexavalents in spermatocytes of Robertsonian heterozygotes between Mus m. domesticus 2n 26 from the Vulcano and Lipari Islands (Aeolian Archipelago, Italy). Eur J Histochem. 2018; 62: 2894. [CrossRef]
- Lyapunova EA, Bakloushinskaya IYu, Kolomiets OL, Mazurova TF. Analysis of fertility of hybrids of multi chromosomal forms in mole voles of the superspecies Ellobius tancrei differing in a single pair of Robertsonian metacentrics. P USSR Acad Sci. 1990; 310: 721–723. (In Russian)
- Matveevsky S, Bakloushinskaya I, Tambovtseva V, Romanenko S, Kolomiets O. Analysis of meiotic chromosome structure and behavior in Robertsonian heterozygotes of Ellobius tancrei (Rodentia, Cricetidae): a case of monobrachial homology. Comp Cytogenet. 2015; 9: 691-706. [CrossRef]
- Bolcun-Filas E, Handel MA. Meiosis: the chromosomal foundation of reproduction. Biol Reprod. 2018. ioy021 doi:10.1093/biolre/ioy021. [CrossRef]
- Kim S, Peterson SE, Jasin M, Keeney S. Mechanisms of germ line genome instability. Semin Cell Dev Biol. 2016; 54: 177-187. [CrossRef]